PART.2
There are also the apparent differences in the distribution of de novo mutations between human and chimpanzee male and female germ lines, as well as between different mutation types. Regarding the human/chimpanzee differences, Harris notes that this is an observation from the scientific literature. "A recent study at Oxford3 sequenced families of chimps and found that many more mutations were transmitted from fathers to children than from mothers to children. This makes sense given that sperm undergo many more cell divisions than do eggs, and thus accumulate mutations faster. This same trend is thought to hold in humans – but in humans, the difference between male and female mutation rates appears less extreme."
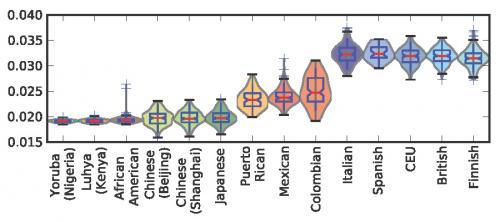
Variation of f(TCC) within and between populations. This plot shows the distribution of f(TCC) within each 1000 Genomes population (i.e., the proportion of all derived variants from PA, PE, and PAf present in a particular genome that are TCC→T mutations). There is a clear division between the low f(TCC) values of African and Asian genomes and the high f(TCC) values of European genomes. The slightly admixed African Americans and more strongly admixed Latin American populations have intermediate f(TCC) values reflecting partial European ancestry. Credit: Harris K (2015) Evidence for recent, population-specific evolution of the human mutation rate. Proc Natl Acad Sci USA 112:11 3439-3444.
Harris also refers to the Oxford study when discussing differences in the distribution of de novo mutations between different mutation types, pointing out that a high percentage of the mutations they found in the chimp family they sequenced were so-called CpG mutations – that is, a mutation in which a C nucleotide separated by a phosphate (p) from, and located next to, a G nucleotide, to a T nucleotide. "This mutation type has a different chemical mechanism than do most mutations where a C changes to a T," Harris explains. "In most organisms, C nucleotides located next to G nucleotides have higher mutation rates than most sites in the genome, and this difference in mutability looks like it might be more extreme within chimp genomes than within human genomes. This is an example of a scenario where one mutation type – in this case, CG->TG – might have increased in rate along the chimp lineage relative to the rates of other mutation types."
Harris has several plans for her continued and future research, one being to analyze whether the mutagenic spectrum might have changed numerous times during great ape evolution and beforehand given that it was able to change during the last 60,000 years of human history. "This is a question I hope to address by applying the techniques I developed in this paper to new datasets. For example," she illustrates, "a large dataset of great ape genomes was published last year, and I'm looking forward to searching for differences between the mutation spectra of great ape species the same way I looked for differences between human populations for this paper. This should give us a better idea of whether the observed difference between Europeans and Africans/Asians was a real anomaly that had functional significance, or whether it was just one of many random mutation rate changes that are more frequent than we've assumed them to be." Harris also plans to investigate DNA from anatomically modern humans who died thousands of years ago, since an increasing number of these ancient DNA sequences are starting to be produced at high quality and released for public use.
Another goal in Harris' calendar is pinpointing finer-scale mutation rate changes and other avenues of research. "The statistical analysis techniques I used in this paper are really simple – I used the same chi-square test taught in high school biology class because the signal of TCC->TTC mutations being more frequent in Europeans than in other groups is so strong that more sophisticated techniques weren't needed to detect it. However, now that I know that the mutational process can differ between human populations, I'm going to develop more robust statistical tools that will be intended to have higher power to detect interpopulation differentiation signals that might be weaker than the dominant signal I studied in this paper, but still be real and thereby important." A key question she'd like to answer is how frequently and routinely mutational processes do change – and she sees detecting smaller differences between populations as important step toward achieving that.
In addition, Harris in interested in determining whether the TCC->TTC excess in Europeans is a trait with a simple genetic basis – that is, if it might be the result of a causal mutation in a single gene associated with DNA replication or repair. "Scientists have mapped the locations of a lot of genes that are associated with variation in traits like height or disease risk, and knowing the genetic basis of the variation in some trait can help you test whether or not that variation was likely driven by natural selection," Harris tells Phys.org. "Usually, we think about a person's DNA sequence encoding a trait that's external to the DNA – but my question is unusual in that I'm interested in a trait that is itself a feature of the DNA sequence." Harris acknowledges that this intertwining presents some technical challenges, but states that if she can develop new techniques for overcoming those challenges, those techniques could allow researchers to study variations in other traits that are endogenous, rather than exogenous, to DNA.
n closing, Harris notes that other areas of research might benefit from her study. "Cancer genetics is an area that I've recently started reading about because of the association of melanoma with the mutation type TCC->TTC that jumped out in the results of my study. Although I'm far from being an expert on cancer genetics at this point, I'm hopeful that that field could have something to gain from a better description of how mutational processes can vary in a benign way between human populations."
More information: Evidence for recent, population-specific evolution of the human mutation rate, Proceedings of the National Academy of Sciences (2015) 112:11 3439-3444, doi:10.1073/pnas.1418652112
Related:
1Genomic data support the hominoid slowdown and an Early Oligocene estimate for the hominoid–cercopithecoid divergence, Proceedings of the National Academy of Sciences (2004) 101(49):17021–17026, 10.1073/pnas.0407270101
2The molecular clock runs more slowly in man than in apes and monkeys, Nature (1987) 326:93-96, doi:10.1038/326093a0
3Strong male bias drives germline mutation in chimpanzees. Science (2014) 344(6189):1272–1275, doi:10.1126/science.344.6189.1272