ARCHEOLOG-HOME / INDIana-UNIversitas
How wild rabbits were genetically transformed into tame rabbits
Uppsala University
Source - http://www.sciencedaily.com/releases/2014/08/140828142744.htm
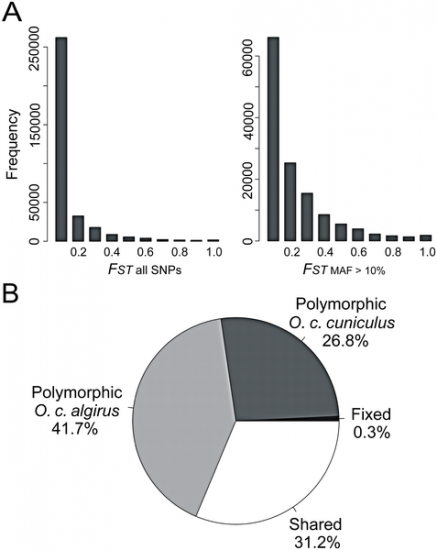
FIGURE 1: Low to moderate overall genetic differentiation between the two subspecies of rabbits.
A) Histogram of FST values for all SNPs and SNPs showing a minor allele frequency (MAF) of 10%. B) Pie chart summarizing the proportion of fixed, shared and exclusive polymorphisms.
doi:10.1371/journal.pgen.1003519.g001
The genetic changes that transformed wild animals into domesticated forms have long been a mystery. An international team of scientists has now made a breakthrough by showing that many genes controlling the development of the brain and the nervous system were particularly important for rabbit domestication. The study is published today in Scienceand gives answers to many genetic questions.
The domestication of animals and plants, a prerequisite for the development of agriculture, is one of the most important technological revolutions during human history. Domestication of animals started as early as 9,000 to 15,000 years ago and initially involved dogs, cattle, sheep, goats and pigs. The rabbit was domesticated much later, about 1,400 years ago, at monasteries in southern France. It has been claimed that rabbits were domesticated because the Catholic Church had declared that young rabbits were not considered meat, but fish, and could therefore be eaten during lent! When domestication occurred, the wild ancestor, the European rabbit (Oryctolagus cuniculus), was confined to the Iberian Peninsula and southern France.
There are several reasons why the rabbit is an outstanding model for genetic studies of domestication: its domestication was relatively recent, we know where it happened, and this region is still densely populated with wild rabbits, explains Miguel Carneiro, from CIBIO/Inbio-University of Porto, one of the leading authors on the paper. Wild rabbits also serve as an excellent model for genetic studies of the early stages of species formation, as shown in an accompanying study we publish today in PLoS Genetics, adds Miguel Carneiro.
The scientists first sequenced the entire genome of one domestic rabbit to develop a reference genome assembly. Then they resequenced entire genomes of domestic rabbits representing six different breeds and wild rabbits sampled at 14 different places across the Iberian Peninsula and southern France.
No previous study on animal domestication has involved such a careful examination of genetic variation in the wild ancestral species. This allowed us to pinpoint the genetic changes that have occurred during rabbit domestication, says Leif Andersson, Uppsala University, Swedish University of Agricultural Sciences and Texas A&M University.
In contrast to domestic rabbits, wild rabbits have a very strong flight response because they are hunted by eagles, hawks, foxes and humans, and therefore must be very alert and reactive to survive in the wild. In fact, Charles Darwin wrote in On the Origin of Species that "…no animal is more difficult to tame than the young of the wild rabbit; scarcely any animal is tamer than the young of the tame rabbit." Darwin used domestic animals as a proof-of-principle that it is possible to change phenotypes by selection. The scientists involved in the current study have now been able to reveal the genetic basis for this remarkable change in behaviour and the study has given important new insights about the domestication process.
Rabbit domestication has primarily occurred by altering the frequencies of gene variants that were already present in the wild ancestor. Our data shows that domestication primarily involved small changes in many genes and not drastic changes in a few genes, states Kerstin Lindblad-Toh, co-senior author, Director of Vertebrate Genome Biology at the Broad Institute of MIT and Harvard, professor at Uppsala University and Co-Director of Science for Life Laboratory.
The team observed very few examples where a gene variant common in domestic rabbits had completely replaced the gene variant present in wild rabbits; it was rather shifts in frequencies of those variants that were favoured in domestic rabbits.
An interesting consequence of this is that if you release domestic rabbits into the wild, there is an opportunity for back selection at those genes that have been altered during domestication because the 'wild-type' variant has rarely been completely lost. In fact, this is what we plan to study next, comments Miguel Carneiro.
The scientists found no example where a gene has been inactivated during rabbit domestication and there were many more changes in the non-coding part of the genome than in the parts of the genome that codes for protein.
The results we have are very clear; the difference between a wild and a tame rabbit is not which genes they carry but how their genes are regulated i. e. when and how much of each gene is used in different cells, explains Miguel Carneiro.
The study also revealed which genes that have been altered during domestication. The researchers were amazed by the strong enrichment of genes involved in the development of the brain and the nervous system, among the genes particularly targeted during domestication.
But that of course makes perfect sense in relation to the drastic changes in behaviour between wild and domestic rabbits, concludes Kerstin Lindblad-Toh.
The study shows that the wild rabbit is a highly polymorphic species that carries gene variants that were favourable during domestication, and that the accumulation of many small changes led to the inhibition of the strong flight response -- one of the most prominent phenotypic changes in the evolution of the domestic rabbit
We predict that a similar process has occurred in other domestic animals and that we will not find a few specific "domestication genes" that were critical for domestication. It is very likely that a similar diversity of gene variants affecting the brain and the nervous system occurs in the human population and that contributes to differences in personality and behaviour, says Leif Andersson.
Journal References:
-
Miguel Carneiro, Frank W. Albert, Sandra Afonso, Ricardo J. Pereira, Hernan Burbano, Rita Campos, José Melo-Ferreira, Jose A. Blanco-Aguiar, Rafael Villafuerte, Michael W. Nachman, Jeffrey M. Good, Nuno Ferrand. The Genomic Architecture of Population Divergence between Subspecies of the European Rabbit. PLoS Genetics, 2014; 10 (8): e1003519 DOI:10.1371/journal.pgen.1003519
-
M. Carneiro, C.-J. Rubin, F. Di Palma, F. W. Albert, J. Alfoldi, A. M. Barrio, G. Pielberg, N. Rafati, S. Sayyab, J. Turner-Maier, S. Younis, S. Afonso, B. Aken, J. M. Alves, D. Barrell, G. Bolet, S. Boucher, H. A. Burbano, R. Campos, J. L. Chang, V. Duranthon, L. Fontanesi, H. Garreau, D. Heiman, J. Johnson, R. G. Mage, Z. Peng, G. Queney, C. Rogel-Gaillard, M. Ruffier, S. Searle, R. Villafuerte, A. Xiong, S. Young, K. Forsberg-Nilsson, J. M. Good, E. S. Lander, N. Ferrand, K. Lindblad-Toh, L. Andersson. Rabbit genome analysis reveals a polygenic basis for phenotypic change during domestication. Science, 2014; 345 (6200): 1074 DOI: 10.1126/science.1253714






















