ARCHEOLOG-HOME / INDIana-UNIversitas
Human genome was shaped by an evolutionary arms race with itself
University of California - Santa Cruz
Source - http://phys.org/news/2014-09-human-genome-evolutionary-arms.html
New findings by scientists at the University of California, Santa Cruz, suggest that an evolutionary arms race between rival elements within the genomes of primates drove the evolution of complex regulatory networks that orchestrate the activity of genes in every cell of our bodies.
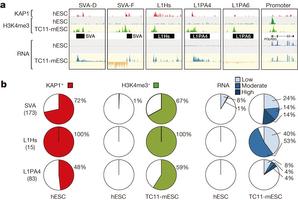
Figure 1: SVAs and L1PAs are derepressed in a non-primate cellular environment.a, KAP1, H3K4me3 ChIP-seq and RNA sequencing (RNA-seq) coverage tracks for a selection of KAP1-bound primate-specific retrotransposons derepressed in TC11-mESCs (yellow) relative to hESCs (grey). H3K4me3 signal on promoters is similar…
The arms race is between mobile DNA sequences known as "retrotransposons" (a.k.a. "jumping genes") and the genes that have evolved to control them. The UC Santa Cruz researchers have, for the first time, identified genes in humans that make repressor proteins to shut down specific jumping genes. The researchers also traced the rapid evolution of the repressor genes in the primate lineage.
Their findings, published September 28 in Nature, show that over evolutionary time, primate genomes have undergone repeated episodes in which mutations in jumping genes allowed them to escape repression, which drove the evolution of new repressor genes, and so on. Furthermore, their findings suggest that repressor genes that originally evolved to shut down jumping genes have since come to play other regulatory roles in the genome.
"We have basically the same 20,000 protein-coding genes as a frog, yet our genome is much more complicated, with more layers of gene regulation. This study helps explain how that came about," said Sofie Salama, a research associate at the UC Santa Cruz Genomics Institute who led the study.
Retrotransposons are thought to be remnants of ancient viruses that infected early animals and inserted their genes into the genome long before humans evolved. Now they can only replicate themselves within the genome. Depending on where a new copy gets inserted into the genome, a jumping event can disrupt normal genes and cause disease. Often the effect is neutral, simply adding to the overall size of the genome. Very rarely the effect might be advantageous, because the added DNA can itself be a source of new regulatory elements that enhance gene expression. But the high probability of deleterious effects means natural selection favors the evolution of mechanisms to prevent jumping events.
Scientists estimate that jumping genes or "transposable elements" account for at least 50 percent of the human genome, and retrotransposons are by far the most common type.
"There have been successive waves of retrotransposon activity in primate evolution, when a transposable element changed to become expressed and replicated itself throughout the genome until something turned it off," Salama said. "We've discovered a major mechanism by which the genome is able to shut down these mobile DNA elements."
The repressors identified in the new study belong to a large family of proteins known as "KRAB zinc finger proteins." These are DNA-binding proteins that repress gene activity, and they constitute the largest family of gene-regulating proteins in mammals. The human genome has over 400 genes for KRAB zinc finger proteins, and about 170 of them have emerged since primates diverged from other mammals.
According to Salama, her team's findings support the idea that expansion of this family of repressor genes occurred in response to waves of retrotransposon activity. Because repression of a jumping gene also affects genes located near it on the chromosome, the researchers suspect that these repressors have been co-opted for other gene-regulatory functions, and that those other functions have persisted and evolved long after the jumping genes the repressors originally turned off have degraded due to the accumulation of random mutations.
"The way this type of repressor works, part of it binds to a specific DNA sequence and part of it binds other proteins to recruit a whole complex of proteins that creates a repressive landscape in the genome. This affects other nearby genes, so now you have a potential new layer of regulation available for further evolution," Salama said.
KRAB zinc finger proteins are the subject of intensive research as scientists try to sort out their many regulatory roles within the genome. The idea that they are involved in repression of jumping genes is not new—previous studies by other researchers have shown that these proteins silence jumping genes in mouse embryonic stem cells. But until now, no one had been able to demonstrate that the same thing occurs in human cells.
The UC Santa Cruz team developed a novel assay to test whether a particular KRAB zinc finger protein could shut down certain jumping genes. The first authors of the paper, postdoctoral researcher Frank Jacobs and graduate student David Greenberg, came up with the strategy of testing primate retrotransposons in non-primate cells by using mouse embryonic stem cells that contain a single human chromosome. In the environment of a mouse cell, jumping genes that were repressed in primate cells became active. Greenberg then developed an assay for testing individual zinc finger proteins for their ability to turn off a primate jumping gene in the mouse cell environment.
"We did all our tests in mouse cells because they lack all of the primate zinc finger proteins, so when you put primate retrotransposons into a mouse cell they're all active," Salama explained.
The results demonstrated that two human proteins called ZNF91 and ZNF93 bind and repress two major classes of retrotransposons (known as SVA and L1PA) that are currently or recently active in primates. Assistant research scientist Benedict Paten directed graduate student Ngan Nguyen in a painstaking analysis of primate genomes, including the reconstruction of ancestral genomes, which showed that ZNF91 underwent structural changes 8 to 12 million years ago that enabled it to repress SVA elements.
Experiments with ZNF 93, which shuts down L1PA retrotransposons, provided a striking illustration of the arms race between jumping genes and repressors. The researchers found that, while it is good at shutting down many L1PA elements, there is one subset of a recently evolved lineage of L1PA that has lost a short section of DNA that includes the ZNF93 binding site. Without the binding site, these jumping genes evade repression by ZNF93. Interestingly, when the researchers put the missing sequence back into one of these genes and put it in a mouse cell without ZNF93, they found that it was better at jumping. So even though the sequence helps with jumping activity, losing it gives the jumping gene an advantage in primates by allowing it to escape repression by ZNF93.
"That's kind of the icing on the cake for aficionados of molecular evolution, because it demonstrates that this is a never-ending race," Salama said. "KRAB zinc finger proteins are a rare class of proteins that is rapidly expanding and evolving in mammalian genomes, which makes sense because the transposable elements are themselves continually evolving to escape repression."
Corresponding author David Haussler, professor of biomolecular engineering and director of the UC Santa Cruz Genomics Institute, said the study involved close collaboration between his group's "wet lab," directed by Salama, and the "dry lab" where researchers under Paten's direction used the computational tools of genome bioinformatics to reconstruct the evolutionary history of primate genomes. Haussler, a Howard Hughes Medical Institute investigator who has used his background in computer science to do pioneering work in genomics, said he established the wet lab to enable just this kind of collaboration.
"Both parts were integral to this study, and there was a lot of back and forth between them. This paper shows how important it is to integrate computational and experimental approaches to fundamental scientific problems, such as how and why we continuously evolve to be more complex," Haussler said.
More information: Nature DOI: 10.1038/nature13760






















